Early administration of appropriate empirical treatment for bloodstream infections (BSI) improves clinical outcomes.1–3 Nevertheless, empiric broad-spectrum antibiotic therapy is often inadequate because the frequent involvement of multiresistant bacteria, among which third-generation cephalosporin-resistant Enterobacterales are of special concern due to their association with increased mortality.4 Matrix-assisted laser desorption/ionization mass spectrometry (MALDI-TOF MS)-based assays provide reliable and timely information on bacterial susceptibility/resistance to antimicrobials.5 Unfortunately, MALDI-TOF MS instruments are not available in all microbiology laboratories. Herein, we developed a simple turbidimetry-based method for detecting resistance to ceftriaxone (CRO) in Escherichia coli and Klebisella spp. from positive blood cultures (BC), which circumvents the use of MALDI-TOF MS instruments. A total of 112 consecutive patients (mean age 74 years; range 40–99; 62.5% male) admitted between November 2019 and June 2020 with BSI due to E. coli (n=77) or Klebsiella spp. (n=35; K. pneumoniae, n=32; K. oxytoca, n=2; K. variicola, n=1) were included. BC bottles were incubated in the Bactec FX automated system (Becton Dickinson, New Jersey, USA). Direct identification of bacteria from BC was carried out using a MALDI-TOF MS protocol6 (Supplementary material). Antimicrobial susceptibility testing (AST) was performed using the broth microdilution MicroScan NM44 panel (Beckman Coulter) and interpreted according to EUCAST guidelines.7 A total of 20 (26%) E. coli and 10 (28.5%) Klebsiella spp. (in all cases K. pneumoniae) isolates were resistant to CRO. Antimicrobial resistance gene characterization of all CRO resistant isolates was carried out using the Antimicrobial Resistance (AMR) Direct Flow Chip (Máster Diagnóstica, Granada, Spain)8 (Supplementary material). All CRO-resistant E. coli isolates and 9/10 CRO-resistant K. pneumoniae harboured a CTX-M type ESBL (extended spectrum beta-lactamase); in turn, 2/10 K. pneumoniae isolated harboured an OXA-48 type D carbapenemase. One CRO-resistant K. pneumoniae presumably harboured a plasmid-mediated AmpC according to the antimicrobial susceptibility profile (piperacillin–tazobactam, MIC ≤8mg/L; cefepime, MIC ≤1mg/L; CRO, MIC >32mg/L; ertapenem, MIC ≤0.5mg/L) and the lack of detection of any beta-lactamase resistance gene by the AMR chip assay.
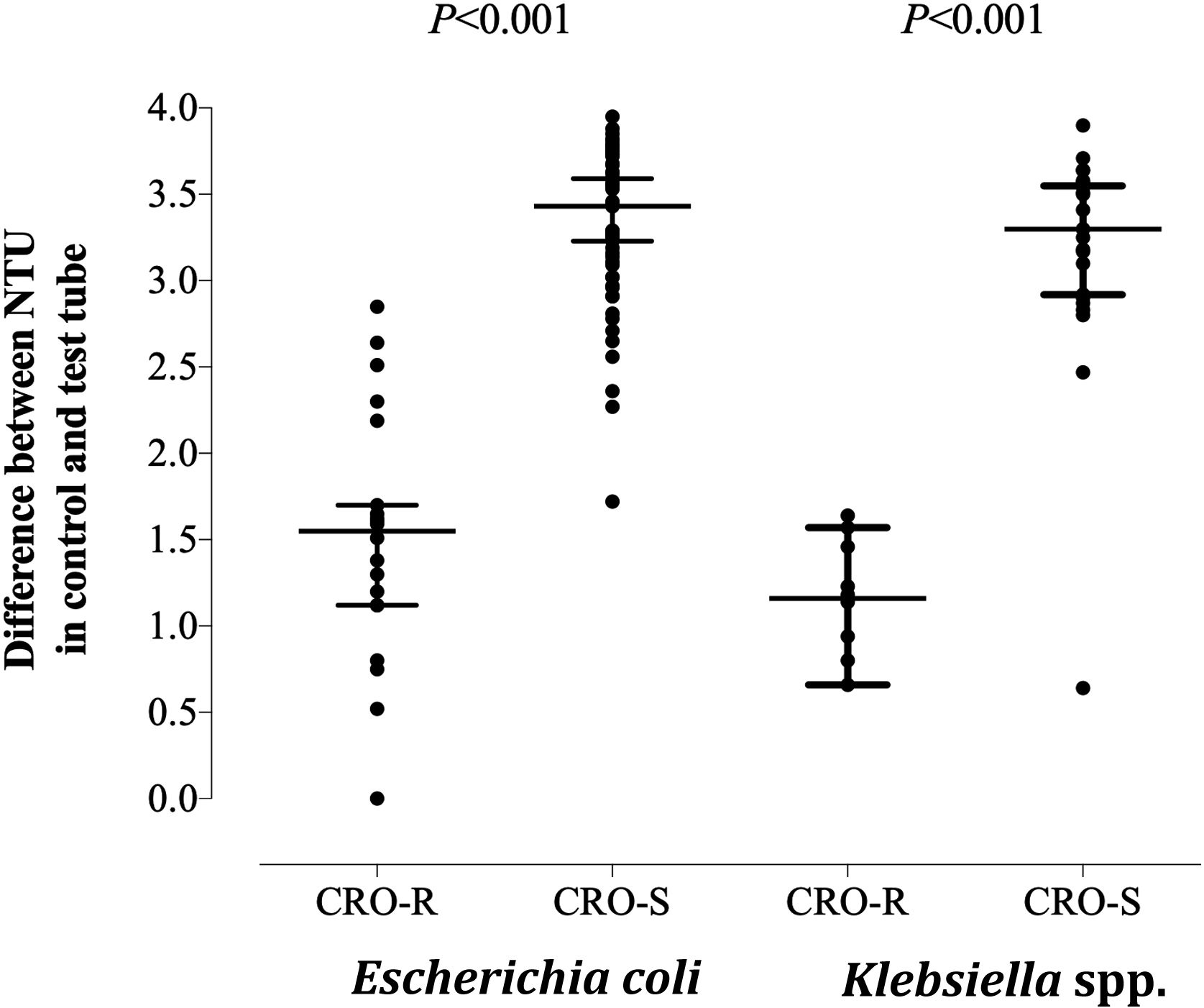
The turbidimetry-based method for detection of CRO resistance in E. coli and Klebsiella spp. (Supplementary material) was conducted as follows. A volume of 50μL of positive BC was incubated with 450μL of brain heart infusion (BHI) broth (Oxoid Limited, Hampshire, UK) in the absence (control) and presence (test) of CRO at a final concentration of 2mg/mL,8 at 37°C in an atmosphere with 5% of CO2 (Heracell 240i CO2 incubator, Thermo Fisher Scientific, Langenselbold, Germany) for 2h. Next, control and test tubes were centrifuged for 3min at 13,000rpm and the resulting pellets were resuspended with 1mL of sterile H2O for subsequent turbidity measurement (Densicheck Plus Instrument, bioMerieux Inc., France). Median nephelometric turbidity units (NTU) value for the control tubes was 3.77 (range 0.86–4), compared with 0.27 (range 0–2.54) and 1.73 (0.27–4) [P<0.001; Mann–Whitney U-test] for the test tubes corresponding to CRO-susceptible and CRO-resistant isolates, respectively. Differential NTU values resulting from the subtraction of NTU values in test tubes from that in control tubes for CRO-susceptible and CRO-resistant E. coli and Klebsiella spp. isolates are shown in Fig. 1. Receiver operating characteristic (ROC) curves determined that overall, a differential NTU value of 2.64 best discriminated (area under the curve of 0.93; P<0.001) between CRO susceptible and resistant isolates, yielding a sensitivity of 96.7% (95% CI, 83.3–99.4; P<0.001) a specificity of 88%, a positive predictive value of 74% and a negative predictive value of 99%; it correctly categorized 29 (96.7%) and 75 (91.5%) of CRO-resistant and CRO-susceptible isolates, respectively. One CTX-M type ESBL-producing E. coli was erroneously categorized as susceptible. As for CRO-susceptible isolates (n=80), 75 (91.5%) were categorized as such by the turbidimetry-based method and 7 were misclassified as resistant. The Kappa correlation index between results obtained by the turbidimetry method and conventional microdilution AST was 0.829 (95% CI, 0.71–0.94). The feasibility of turbidimetry to reliable predict antimicrobial susceptibility directly from bacteria in grown BC was shown for the Alfred AST® system (Alifax, Padova, Italy).9 In summary, the turbidimetry-based method described herein reliably predicts CRO resistance in E. coli and Klebsiella spp.; the method is cheap and accessible to all clinical microbiology laboratories. Nevertheless, due to the limited number of CRO-resistant isolates included, its validation requires further studies.
Differential nephelometric turbidity units (NTU) values resulting from the subtraction of NTU values in test tubes (with ceftriaxone at 2mg/mL) from that in control tubes (without ceftriaxone-CRO-) for CRO-susceptible and CRO-resistant E. coli and Klebsiella spp. recovered from blood cultures. P values for comparisons (Mann–Whitney U test) are shown. Statistical significance was set at P value <0.05.
DS and IT: methodology and data validation; DS, IT, JC and DN: study design and logistics; DN and IT: conceptualization, data analysis and manuscript writing.
FundingThe authors declare that no funds, grants, or other support were received during the preparation of this manuscript.
Ethical approvalThe current study was approved by the Research Ethics Committee of Hospital Clínico Universitario INCLIVA (September, 2019).
Conflict of interestThe authors have no conflict of interest.
We are grateful to residents and staff at the Microbiology Service of Hospital Clínico Universitario. Ignacio Torres holds a contract (Río Hortega Contract; CM20/00090) funded by the Carlos III Health Institute (co-financed by the European Regional Development Fund, ERDF/FEDER).